What is phylogeny?Phylogeny is the study of ancestral linkages and divergences over a period of time and helps to determine the relatedness of different species based on their genes, proteins, or physical features. The trees below depict various types of ways to look at how all of the genes can be arranged to show similarities [1].
All of the following trees were created on Clustal Omega. |
Blosum matrixTwo of the trees to the right describe the methods that would be achieved by using a BLOSUM62 Matrix. BLOSUM62 is a scoring system used to calculate the similarity between sequences. BLOSUM scores the data between two protein sequences and aligns them based on their amino acids. It is determined by whether or not the sequences is the same and how likely that is to be random. A total score is then achieved and we can see how related they are to each other.
Percent IdentityPercent identity is fairly self explanatory in that it shows the percentage of how identical the sequences are. You can used neighbor going and average distance as methods using this.
Neighbor JoiningNeighbor joining is a secondary step to the two above in that it takes the data that has been calculated by the percent identity and BLOSUM and then uses both to see which ones are more related.
Average DistanceAverage distance also uses the Percent Identity and BLOSUM62 scores to determine which species are more similar and then rates them based on the length. This would show that each organism has diverged from the same common ancestor.
|
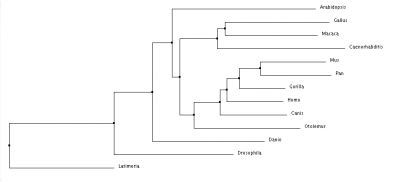
Neighbor joining tree using BLOSUM62
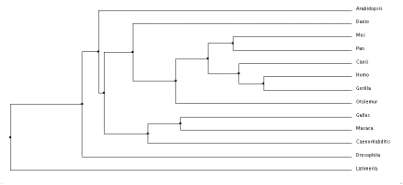
Average distance tree using BLOSUM62
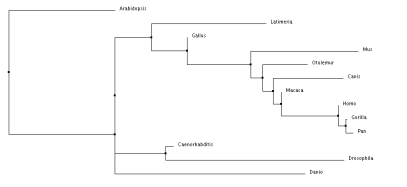
Neighbor Joining tree using PID
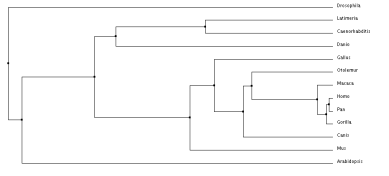
Average distance tree using PID
|
Discussion
After comparing all of the trees there is a lot of similarities between the groups and generally what changes has to do with the outlier of the group, occasionally it is A. thaliana which is understandable, because it is the only plant in the matrix. But it also occasionally has the coelacanth on the outskirts which is odd that it is not always along with the zebrafish, as all the primates are together but that explains how diverse the fish population is. As expected, the primates are all grouped together as this is a skin cell-based disease and we should see the most genetic similarity in organisms that could potentially experience the same fate.
References:
[1] Understanding Phylogenies (1 of 2). (n.d.). Retrieved March 28, 2015, from http://evolution.berkeley.edu/evosite/evo101/IIBPhylogenies.shtml
[2] Where did the BLOSUM62 alignment score matrix come from? Sean R Eddy
[3] BLAST Glossary: http://www.ncbi.nlm.nih.gov/books/NBK62051/
[4] Evolution textbook: http://evolution-textbook.org/content/free/contents/ch27.html#ch27-4-2
[1] Understanding Phylogenies (1 of 2). (n.d.). Retrieved March 28, 2015, from http://evolution.berkeley.edu/evosite/evo101/IIBPhylogenies.shtml
[2] Where did the BLOSUM62 alignment score matrix come from? Sean R Eddy
[3] BLAST Glossary: http://www.ncbi.nlm.nih.gov/books/NBK62051/
[4] Evolution textbook: http://evolution-textbook.org/content/free/contents/ch27.html#ch27-4-2