Identification of Phosphorylation sites on Par-5 and its binding partners.
Introduction:
Colorectal cancer is third leading cause of cancer deaths in the United States and it is comprised of colon and rectal cancers. It is an invasive disease that does not tend to show many warning signs until the disease has progressed into a stage 4 or 5 in which it is almost possible for survival [1]. In 2015 alone, there are an expected 50,310 deaths and 136,830 new cases in the United States. This type of cancer in particular affects the intestinal epithelial cells or the keratinocytes in the colon, there is an inability to suppress growing tumors and malignant polyps form [1]. This is usually followed by visible and physical symptoms of stomach cramps, vomiting, nausea and blood in the stool. These are signs that insinuate that there are bleeding polyps forming in the colon, however many people assume they have a bad case of the flu until the bloody stool appears [1]. This can be caused by a defect in any number of genes, since cancer is a polygenic disorder and can be affected by genetics and environmental factors. However, one gene in particular, 14-3-3σ is a known tumor suppressor gene and it is found to be down-regulated in patients with different types of cancer [2]. It is also very well conserved across vertebrates, invertebrates and plants [3].
The primary goal research presented below will delve into understanding the role that 14-3-3σ plays in cell division and how its depletion leads to tumorigenesis. It is a negative regulator of p53 and cyclin dependent kinases, it also plays a role in mitotic cell arrest in order to repair DNA damage, it also plays a specific role in translation. Mutations in 14-3-3σ also lead to cytokinesis failures [4]. While much is known about the function of 14-3-3sigma in translation, it is still unclear is how 14-3-3σ regulates cell division and which 14-3-3σ binding partners aid in this cellular process. My hypothesis is that 14-3-3σ/PAR-5 and its binding partners regulate the activity proteins necessary for cytokinesis by binding to serine/threonine sites. In order to achieve a better understanding of how this protein is working in the cell, I will be using the C. elegans 14-3-3σ homolog, PAR-5, because cell division is easily assayed and imaged in C. elegans embryos [6]. When this has been negated or accepted, the long-term goal is to determine how 14-3-3σ functions in cell division events. The following aims describe how this hypothesis will be tested.
Aim 1: The main purpose of this aim is to identify and characterize the function of 14-3-3σ/PAR-5 interacting partners and determine if they play a role in cell division. To identify these interacting partners I used STRING data to identify binding partners of PAR-5 in C. elegans and 14-3-3σ to see if there was any overlap between the interactions. The data from this experiment can be seen here.
Then using Gene Ontology, I sorted the proteins based on biological function to determine if these interactions play a role in cell division. This step was actually simplified by the fact that there was only one protein that overlapped homologs between the protein-protein interaction network that was hopeful to have a function in phosphorylation sites and cell division, which was sir-2.1 (SIRT1: human homolog). Sir-2.1 is a NAD-dependent deacetylase SIR2 homolog ; Functions upstream of daf-16 in the insulin-like signaling pathway, promoting daf-16 mediated transcriptional activation and increased life-span. May also regulate life-span independently of daf-16 by modulating the transcription of genes involved in the stress response of the endoplasmic reticulum [4]. Then I used SMART to see if they share similar domains known to be regulated during cell division. These proteins did not share a domain, however I BLASTed them together and they share similar conserved regions. This lead to my next aim as to where they shared conserved regions, if the fell in the active domains and if they shared any motifs.
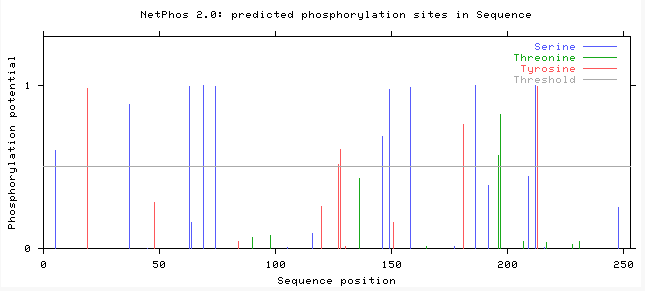
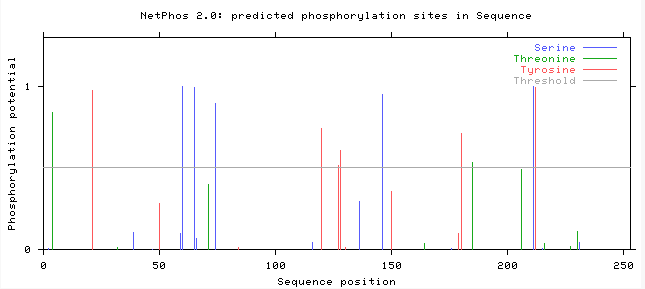
Aim 2: The main purpose of this aim was to identify conserved domains and serine/threonine phosphorylation sites on PAR-5 and Sir-2.1. In order to do this, I performed a NetPhos 2.0 experiment and compared it to the motif data that I found from MEME. NetPhos 2.0 determines where the post-translational phosphorylations occur on the protein, whether they are tyrosine, threonine or serine phosphorylations. Since 14-3-3σ and PAR-5 both function in phospho-threonine and phospho-serine binding, I performed an experiment only using those two. The data I obtained can be seen in the following images:
After viewing this data and comparing how similar the motifs were, I was able to determine the locations that were the most conserved based on the NetPhos 2.0 active sites and the motifs that were similar between the proteins [citation].
For sir-2.1 (amino acid residues 171-220) and par-5 (amino acid residues 119-168) they shared a 50 amino acid residue motif, which fell between the active domains on both proteins. I then compared these sites with the NetPhos 2.0 data, which showed that there was a phosphoserine at amino acid residue 218 on sir-2.1 and one at the amino acid residue 146 on PAR-5 which could mean that this is where they are binding to each other.
To make sure that this interaction does not occur only in C. elegans, I compared this data to the human protein homologs SFN and SIRT1. They also shared a 50 amino acid residue motif in the active domain portion of the protein, which is seen in the above figure. They also shared conserved phosphorylation sites on this domain. Now, in order to determine whether or not these proteins are interacting I will need to perform a knockout assay using CRISPR-Cas9 technology, which leads me to my next aim.
Aim 3: The main purpose of this aim is to determine whether or not sir-2.1 and PAR-5 are working together to mediate cell division and what occurs in the cells when one or both of these proteins are non-functioning. In order to accomplish this, I will use C. elegans and CRISPR-Cas9 technology (which is laid out in more detail here), I will mutate conserved phosphorylation sites by exchanging the amino acids where the phosphorylations sites normally occur to a different amino acid (such as glutamic acid). I will do this in PAR-5, sir-2.1 and potentially other proteins such as lin-45, to create transgenic lines and determine which sites are important for cytokinesis via video microscopy. Once this is determined, I will then tag these proteins with GFP via CRISPR and determine if they localize to cell division structures, such as the spindle or actin ring.
This experiment will have to be carried out by making various transgenic lines which would include an empty vector, PAR-5, PAR-5 and Sir-2.1, Sir-2.1, Lin-45 and one with all three proteins. The vector and PAR-5 lines will be used as a control since research has already been done as to how PAR-5 will react when it is depleted [6]. By performing this assay, I can identify phosphorylation sites on PAR-5 interacting proteins that are important for cytokinesis and then determine if these proteins localize to structures involved during cell division events.
Future Directions:
With this knowledge, I will be able to apply the fundamental aspects of this research to well conserved binding partners of 14-3-3σ to discovery interaction pathway targets in cell division that are necessary for tumor growth. This would allow for treatments/drugs to be created at these sites. If the CRISPR-Cas9 technology confirms my hypothesis then drugs could be created to target those phosphorylation sites, and determine where upstream there are proteins that would cause a down regulation of 14-3-3σ in the cells. By determining these locations when can hope to enhance the tumor suppression function. Millions of people depend on this type of research, not just colorectal cancer patients but also patients that have any 14-3-3 regulated cancer, mostly breast and ovarian cancers [5]. If my hypothesis is negated, then I will go back and look more into other proteins that interact with 14-3-3 proteins, since they seem to be the missing link to cancer research.
For sir-2.1 (amino acid residues 171-220) and par-5 (amino acid residues 119-168) they shared a 50 amino acid residue motif, which fell between the active domains on both proteins. I then compared these sites with the NetPhos 2.0 data, which showed that there was a phosphoserine at amino acid residue 218 on sir-2.1 and one at the amino acid residue 146 on PAR-5 which could mean that this is where they are binding to each other.
To make sure that this interaction does not occur only in C. elegans, I compared this data to the human protein homologs SFN and SIRT1. They also shared a 50 amino acid residue motif in the active domain portion of the protein, which is seen in the above figure. They also shared conserved phosphorylation sites on this domain. Now, in order to determine whether or not these proteins are interacting I will need to perform a knockout assay using CRISPR-Cas9 technology, which leads me to my next aim.
Aim 3: The main purpose of this aim is to determine whether or not sir-2.1 and PAR-5 are working together to mediate cell division and what occurs in the cells when one or both of these proteins are non-functioning. In order to accomplish this, I will use C. elegans and CRISPR-Cas9 technology (which is laid out in more detail here), I will mutate conserved phosphorylation sites by exchanging the amino acids where the phosphorylations sites normally occur to a different amino acid (such as glutamic acid). I will do this in PAR-5, sir-2.1 and potentially other proteins such as lin-45, to create transgenic lines and determine which sites are important for cytokinesis via video microscopy. Once this is determined, I will then tag these proteins with GFP via CRISPR and determine if they localize to cell division structures, such as the spindle or actin ring.
This experiment will have to be carried out by making various transgenic lines which would include an empty vector, PAR-5, PAR-5 and Sir-2.1, Sir-2.1, Lin-45 and one with all three proteins. The vector and PAR-5 lines will be used as a control since research has already been done as to how PAR-5 will react when it is depleted [6]. By performing this assay, I can identify phosphorylation sites on PAR-5 interacting proteins that are important for cytokinesis and then determine if these proteins localize to structures involved during cell division events.
Future Directions:
With this knowledge, I will be able to apply the fundamental aspects of this research to well conserved binding partners of 14-3-3σ to discovery interaction pathway targets in cell division that are necessary for tumor growth. This would allow for treatments/drugs to be created at these sites. If the CRISPR-Cas9 technology confirms my hypothesis then drugs could be created to target those phosphorylation sites, and determine where upstream there are proteins that would cause a down regulation of 14-3-3σ in the cells. By determining these locations when can hope to enhance the tumor suppression function. Millions of people depend on this type of research, not just colorectal cancer patients but also patients that have any 14-3-3 regulated cancer, mostly breast and ovarian cancers [5]. If my hypothesis is negated, then I will go back and look more into other proteins that interact with 14-3-3 proteins, since they seem to be the missing link to cancer research.
| simonsmikaylaupdatedpresentation.pdf | |
| File Size: | 5149 kb |
| File Type: | |
| simonsmikaylaroughdraftpresentation2.pdf | |
| File Size: | 4344 kb |
| File Type: | |
| simonsmikaylaroughdraftpresentation.zip | |
| File Size: | 5514 kb |
| File Type: | zip |
References:
[1] Colon and Rectal Cancer. (n.d.). Retrieved February 19, 2015, from http://www.cancer.gov/cancertopics/types/colon-and-rectal
[2] Yang, H., Wen, Y., Chen, C., Lozano, G., & Lee, M. (2003). 14-3-3 Positively Regulates p53 and Suppresses Tumor Growth. Molecular and Cellular Biology, 7096-7107.
[3] Wilker, E.W., van Vugt, M., Artim, S.A., Huang, P.H., Petersen, C.P., Reinhardt, H.C., Feng, Y., Sharp, P.A., Sonenberg, N., White, F.M., et al. (2007). 14-3-3 sigma controls mitotic translation to facilitate cytokinesis. Nature 446, 329-332.
[4] NAD-dependent protein deacetylase sir-2.1. (n.d.). Retrieved May 18, 2015, from http://www.uniprot.org/uniprot/Q21921
[5] Hermeking, H. (2003). The 14-3-3 cancer connection. Nature Reviews Cancer, 3(12), 931-943. doi: 10.1038/nrc1230.
[6] Morton, D., Shakes, D., Nugent, S., Dichoso, D., Wang, W., Golden, A., & Kemphues, K. (n.d.). The Caenorhabditis elegans par-5 Gene Encodes a 14-3-3 Protein Required for Cellular Asymmetry in the Early Embryo. Developmental Biology, 47-58.
[1] Colon and Rectal Cancer. (n.d.). Retrieved February 19, 2015, from http://www.cancer.gov/cancertopics/types/colon-and-rectal
[2] Yang, H., Wen, Y., Chen, C., Lozano, G., & Lee, M. (2003). 14-3-3 Positively Regulates p53 and Suppresses Tumor Growth. Molecular and Cellular Biology, 7096-7107.
[3] Wilker, E.W., van Vugt, M., Artim, S.A., Huang, P.H., Petersen, C.P., Reinhardt, H.C., Feng, Y., Sharp, P.A., Sonenberg, N., White, F.M., et al. (2007). 14-3-3 sigma controls mitotic translation to facilitate cytokinesis. Nature 446, 329-332.
[4] NAD-dependent protein deacetylase sir-2.1. (n.d.). Retrieved May 18, 2015, from http://www.uniprot.org/uniprot/Q21921
[5] Hermeking, H. (2003). The 14-3-3 cancer connection. Nature Reviews Cancer, 3(12), 931-943. doi: 10.1038/nrc1230.
[6] Morton, D., Shakes, D., Nugent, S., Dichoso, D., Wang, W., Golden, A., & Kemphues, K. (n.d.). The Caenorhabditis elegans par-5 Gene Encodes a 14-3-3 Protein Required for Cellular Asymmetry in the Early Embryo. Developmental Biology, 47-58.