What are interaction Networks?
Interaction networks take the protein-protein interactions described by the interactions of their domains and compare how they all interact in the cell together. It takes one centralized protein and using domain-domain interactions connects the proteins based on their molecular function, cellular components and biological processes, an example can be seen below.
For example, par-5 has been studied extensively so there is more known about its protein interactions within C.elegans and provides a web of the different types of biological processes that could be working together. 14-3-3sigma, which functions with proteins that regulate cell division and translation, and those proteins also function with others that regulate cell division and translation, meaning that there could be interacting with each other to create a problem for the overall cell-cycle process.
After comparing the PAR-5 and 14-3-3sigma interaction networks, I will be looking for proteins that have the most similar GO terms to see how the different modifications of those proteins allow them to act together and see if they have similar domains or binding motifs. In this way, I will be able to see what kind of small molecules could be created to target those sequences, should the interactions fail we can target the domains on multiple proteins in order to rescue the 14-3-3sigma failure.
In this way, we would be able to rescue the tumor suppressor function, and prevent ripple effects from proteins that interact with 14-3-3sigma. This could potentially in an ideal situation be a cure for cancer.
In order to create these charts, I have used STRING, PFAM and SMART to compile the data and then cross-referenced the proteins of interested with BLAST and GO to see the different terms that it showed and compare them from there.
For example, par-5 has been studied extensively so there is more known about its protein interactions within C.elegans and provides a web of the different types of biological processes that could be working together. 14-3-3sigma, which functions with proteins that regulate cell division and translation, and those proteins also function with others that regulate cell division and translation, meaning that there could be interacting with each other to create a problem for the overall cell-cycle process.
After comparing the PAR-5 and 14-3-3sigma interaction networks, I will be looking for proteins that have the most similar GO terms to see how the different modifications of those proteins allow them to act together and see if they have similar domains or binding motifs. In this way, I will be able to see what kind of small molecules could be created to target those sequences, should the interactions fail we can target the domains on multiple proteins in order to rescue the 14-3-3sigma failure.
In this way, we would be able to rescue the tumor suppressor function, and prevent ripple effects from proteins that interact with 14-3-3sigma. This could potentially in an ideal situation be a cure for cancer.
In order to create these charts, I have used STRING, PFAM and SMART to compile the data and then cross-referenced the proteins of interested with BLAST and GO to see the different terms that it showed and compare them from there.
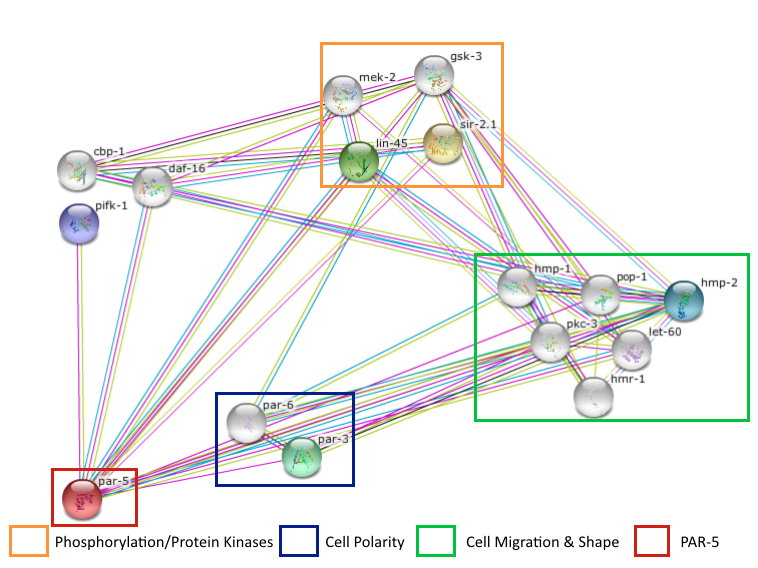
Par-5 Interaction Network
This STRING shows the different interactions with par-5 and multiple GO Functions are compared.
The most note worthy interactions that I will pursue are sir-2.1 (human homolog: sirtuin-1) and lin-45 (human homolog: BRAF)
The most note worthy interactions that I will pursue are sir-2.1 (human homolog: sirtuin-1) and lin-45 (human homolog: BRAF)
- sir-2.1: NAD-dependent deacetylase SIR2 homolog ; Functions upstream of daf-16 in the insulin-like signaling pathway, promoting daf-16 mediated transcriptional activation and increased life-span. May also regulate life-span independently of daf-16 by modulating the transcription of genes involved in the stress response of the endoplasmic reticulum [1].
- lin-45: Raf homolog serine/threonine-protein kinase ; Protein kinase that participates in the induction of vulva and has roles in fertility and viability [1].
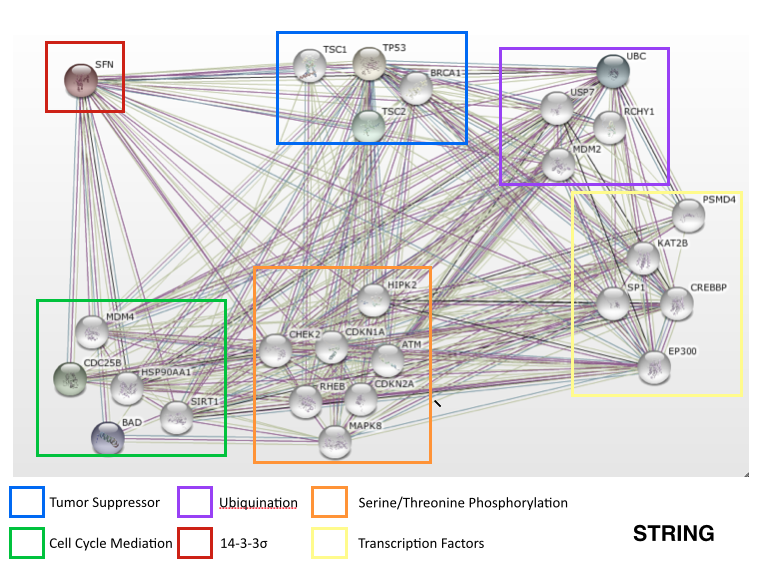
14-3-3sigma interaction network
After reviewing the interaction networks. There was a human homolog for both sir-2.1 and lin-45 that interacted with 14-3-3sigma (SFN) however the only one that showed up on the interaction network was SIRT1, the sir-2.1 human homolog, which in humans aids in cell cycle regulation.
- SIRT1: NAD-dependent protein deacetylase that links transcriptional regulation directly to intracellular energetics and participates in the coordination of several separated cellular functions such as cell cycle, response to DNA damage, metobolism, apoptosis and autophagy [1].
Discussion
The interaction networks confirm the research and that there are many proteins that aid in cell cycle, division and tumor suppressor function, none of which came to my surprise. For the purpose of my hypothesis, it is important to determine what and how some of the other phosphorylation proteins interact with 14-3-3 sigma. It is interesting to note that these interactions do have a homolog between the two interaction networks and that is sir-2.1 and it has to deal with the cell cycle and modifications to transcription genes and they have different functions in each organism. So this is the like the protein I will see having a reaction with the phospho-serine binding that
14-3-3sigma engages in.
14-3-3sigma engages in.